当前位置: 首页 > 生化试剂 > SYBR® Green I Nucleic Acid Gel Stain

描述
SYBR Green I 核酸凝胶染料是极其灵敏的核酸凝胶染料,在结合 dsDNA 时发出明亮的荧光,而且凝胶背景很低,因此对于使用激光扫描仪或标准 UV 透射仪检测凝胶中 dsDNA 的应用来说非常理想。 SYBR Green I 核酸染料也可用于毛细管电泳、实时 PCR 测定和条带偏移测定中。 500 µL 可对 50 块微型凝胶进行染色。 该染料可以 1 ml 单位大小 (S-7567) 和特殊包装 (S-7585) 提供。
规格
Target Molecule: DNA Label or Dye: SYBR® Green I Detection Method: Fluorescent Detection Location: In-Gel Detection Product Size: 500 µL Shipping Condition: Room Temperature
内容及储存
Supplied as a 10,000X concentrate in DMSO. Store at -20°C, protected from light in a dessicatorFigures
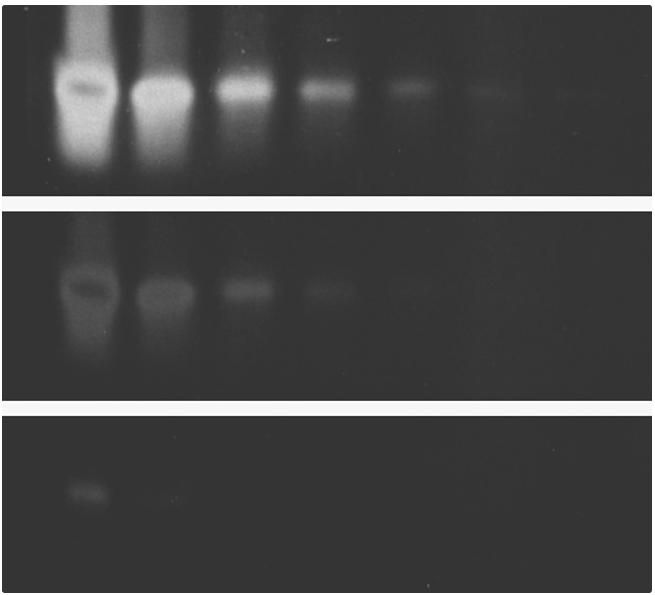
Comparison of single-stranded oligonucleotide detection using SYBR® Green I Nucleic Acid Gel Stain and ethidium bromide.
Identical three-fold dilutions of a synthetic, single-stranded 24-mer were electrophoresed on 10% polyacrylamide gels. Gels were stained for 30 min with a 1:10,000 dilution of SYBR® Green I Nucleic Acid Gel Stain (Cat. No. S7563, S7567, S7585) and not destained (panels A and B) or with 5 µg/mL ethidium bromide (Cat. No. E1305, E3565) for 30 min and destained for a further 30 min in water (panel C). Gel staining was visualized using 254 nm epi-illumination (panel A) or 300 nm transillumination (panels B and C) and then photographed using Polaroid 667 black-and-white print film and a SYBR® photographic filter (Cat. No. S7569, panels A and B) or an ethidium bromide gel stain photographic filter (panel C).
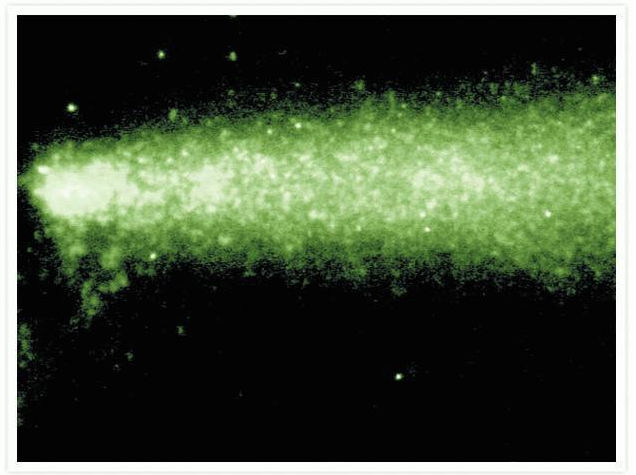
HL-60 cells in a comet assay.
Comet assay with SYBR® Green I Nucleic Acid Gel Stain (Cat. No. S7563, S7567, S7585). DNA fragmentation associated with oxidative DNA damage was visualized using Trevigen's CometAssay kit. HL-60 cells were treated with hydrogen peroxide and immobilized onto a Trevigen CometSlide for analysis. The cells were gently lysed, washed, and treated with endonuclease. Slides were subjected to electrophoresis in alkaline electrophoresis buffer and stained with SYBR® Green I stain.
Agarose gel containing camptothecin-treated HL-60 cells.
DNA extracts from camptothecin-treated HL-60 cells were separated on an agarose gel and stained with SYBR® Green I Nucleic Acid Gel Stain (Cat. No. S7563, S7567, S7585). The 200 to 5,000 bp DNA fragments characteristic of apoptotic cells (which appear as "ladders") are clearly visualized with this sensitive nucleic acid stain. Cell preparations were gifts of Zbigniew Darzynkiewicz, Cancer Research Institute, New York Medical College.
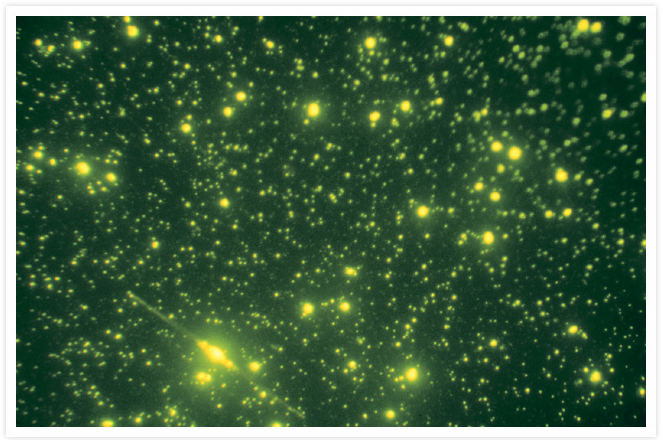
Marine viruses, bacteria, and a diatom.
An environmental sample containing marine viruses (smallest dots), bacteria (larger, brighter dots), and a diatom (long thin cell with prominent nucleus) were stained with SYBR® Green I Nucleic Acid Stain (Cat. No. S7563, S7567, S7585). Image contributed by Jed Fuhrman, University of Southern California.
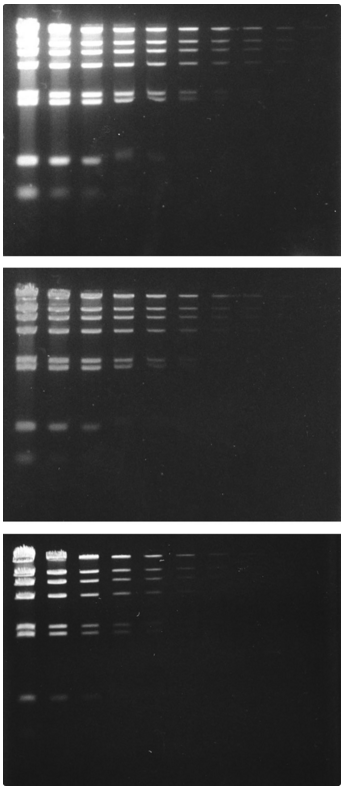
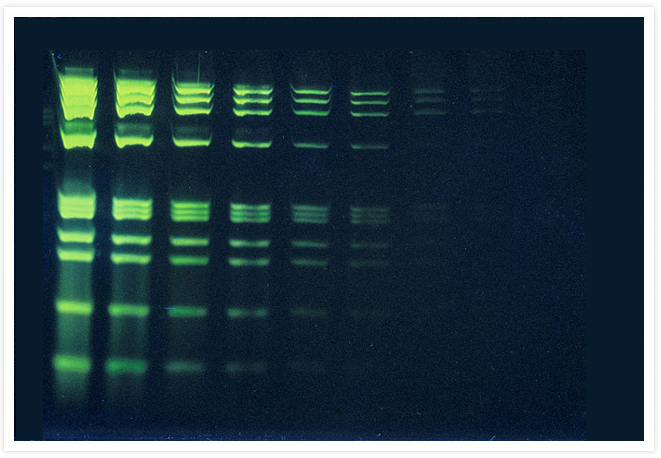
Comparison of dsDNA detection in native gels using SYBR® Green I Nucleic Acid Gel Stain and ethidium bromide.
Identical three-fold dilutions of bacteriophage DNA digested with HindIII restriction endonuclease were electrophoresed on 1% agarose gels. Gels were stained for 30 min with a 1:10,000 dilution of the SYBR® Green I Nucleic Acid Gel Stain (Cat. No. S7563, S7567, S7585) and not destained (panels A and B) or with 5 µg/mL ethidium bromide (Cat. No. E1305, E3565) for 30 min and destained for a further 30 min in water (panel C). Gel staining was visualized using 254 nm epi-illumination (panel A) or 300 nm transillumination (panels B and C) and then photographed using Polaroid 667 black-and-white print film and a SYBR® photographic filter (panels A and B, Cat. No. S7569) or an ethidium bromide gel stain photographic filter (panel C).
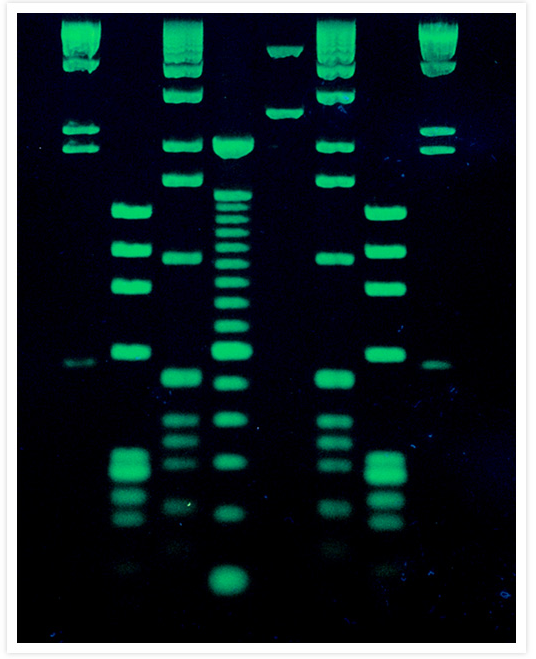
DNA molecular weight ladders stained with SYBR® Green I Nucleic Acid Gel Stain.
DNA molecular weight ladders that were electrophoresed on a 1% agarose gel were stained for 30 min with a 1:10,000 dilution of SYBR® Green I Nucleic Acid Gel Stain (Cat. No. S7563, S7567, S7585). Lanes 1 and 8 contain HindIII–cut DNA; lanes 2 and 7, HaeII–cut fX174 RF DNA; lanes 3 and 6, 1 kilobase pair DNA ladder (Life Technologies); lane 4, 100 Base Pair DNA Ladder (Life Technologies); lane 5, EcoRI–cut pUC19 DNA mixed with PstI–cut fX174 RF DNA. Gel staining was visualized using 254 nm epi-illumination.
Stained HaeIII–digested PhiX174 RF DNA.
A two-fold dilution series of HaeIII–digested PhiX174 RF DNA that was electrophoresed on a 5% polyacrylamide gel was stained with our ultrasensitive SYBR® Green I Nucleic Acid Gel Stain (Cat. No. S7563, S7567, S7585). Gel staining was visualized with 254 nm epi-illumination.
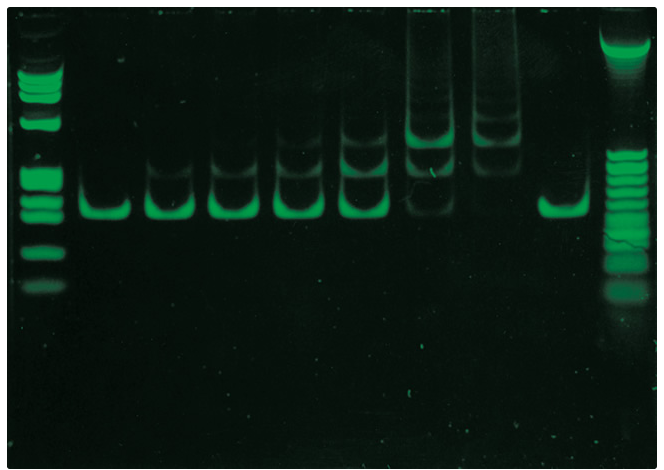
Electrophoretic bandshift assay using SYBR® Green I Nucleic Acid Gel Stain.
The association between a 208-bp fragment purified from AvaI–digested plasmid p5S208-12 and a mutant restriction endonuclease (EcoRI/Gln111) was analyzed on a 4% native polyacrylamide gel. Samples containing approximately 50 ng total fragments and various amounts of the mutant enzyme were subjected to electrophoresis and stained with SYBR® Green I Nucleic Acid Gel Stain. Gel staining was visualized using 254 nm epi-illumination and then photographed using Polaroid 667 black-and-white print film and a SYBR® photographic filter (Cat. No. S7569). Lane 1 contains HaeIII-digested FX174 RF DNA markers; lanes 2 through 9 contain 0, 0.05, 0.1, 0.2, 0.4, 0.6, 0.8 and 0 µM EcoRI/Gln111; lane 10 contains HhaI-digested plasmid p5S208-12 as a size standard.
荧光光谱
SYBR Green
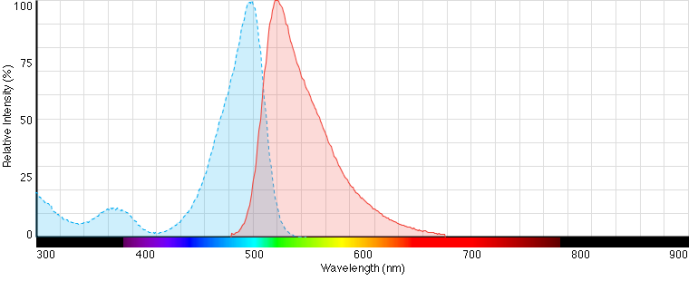
Fluorescence Ex/Em spectra of SYBR® Green I nucleic acid gel stain bound to DNA.











 Global - English
Global - English
 China - 中文
China - 中文
 Brasil - Português
Brasil - Português
 France - Français
France - Français
 Germany - Deutsch
Germany - Deutsch
 Hungary - Magyarország
Hungary - Magyarország
 Italy - Italiano
Italy - Italiano
 Japan -日本语
Japan -日本语
 Russia - Россия
Russia - Россия
 United States - English
United States - English

